Semester Overview, 2022¶

Image credit: Lilian Matallana
Class meetings are in Jordan Hall 6117 from 8:30 to 10:20 am on Mondays, Wednesdays, and Fridays.
The Biostar Handbook is a resource for much of the reading.
A Slack channel is available for questions and discussion.
Will Kohlway (whkohlwa at ncsu.edu) is the teaching assistant for the course.
Course Schedule¶
Week and Dates Topic [Relevant Biostar Handbook chapters, links to other resources]
Week 1, 10 – 14 Jan Introduction to bioinformatics, Linux, and the command-line interface [1, 3, The Linux Command Line, Data Science at the Command Line]
17 Jan Martin Luther King, Jr Day - University closed, no classes
Week 2, 19 - 21 Jan Sequencing instruments [12, Performance Assessment of DNA Sequencing Platforms], experimental design, and introduction to the NC State HPC
Week 3, 24 - 28 Jan Data preprocessing and quality control [14] and sequence read alignment [28]
Week 4, 31 Jan - 4 Feb Transcriptome assembly [25 - general introduction; Best Practices for DeNovo Transcriptome Assembly with Trinity ]
Week 5, 7 - 11 Feb Genome assembly [25]
Week 6, 14 - 18 Feb Re-sequencing, alignment, structural variation [18]
Week 7, 21 - 25 Feb Discovery and genotyping of genetic variation [20, 21]
Week 8, 28 Feb - 4 Mar R and R Studio - Software Carpentry Programming with R and R for Reproducible Scientific Analysis, Data Carpentry Genomics Workshop
Week 9, 7 - 11 Mar Transcriptome analysis: differential gene expression, annotation [22]
Week 10, 14 - 18 Mar Spring Break - no classes
Week 11, 21 - 25 Mar Genome analysis: ChIP-seq, DHS-seq, 3-D conformation [23, 24, CLC GWB ChIP sequencing tutorial]
Week 12, 28 Mar - 1 Apr Linux command-line tools: awk, sed, and bash [Bioawk Basics; Sed Basics; Using bash in bioinformatics; The Art of Bioinformatics Scripting]
Week 13, 4-8 Apr CLC Genomics Workbench - data QC and pre-processing
Week 14, 11 - 15 Apr CLC Genomics Workbench - Expression analysis using RNA-seq data [tutorial]
Week 15, 18 - 22 Apr CLC Genomics Workbench - Paired-end short-read genome assembly [tutorial]; Long-read assembly with short-read polishing [tutorial]
25 Apr Last day of class - questions, review, summary
General background information and course resources¶
- General advice on troubleshooting
- Course syllabus
- Lior Pachter’s list of sequencing-based assays: *Seq
- DNA Sequencing Methods Collection - Illumina compilation of published methods
- For All You Seq - DNA and For All You Seq - RNA are Illumina posters that give brief graphical summaries of many different kinds of sequencing assays
- The R statistical programming environment
- Course resources and recommended reading
- Overview slides (old and outdated in some respects)
- Cloud computing with Amazon Web Services: setting up an AWS account and starting an instance
- OMICtools software database publication and website
- Qiagen webpage of tutorials for CLC Workbench programs
- Colib’read project webpage on reference-assembly-free programs for SNP and indel detection and moress
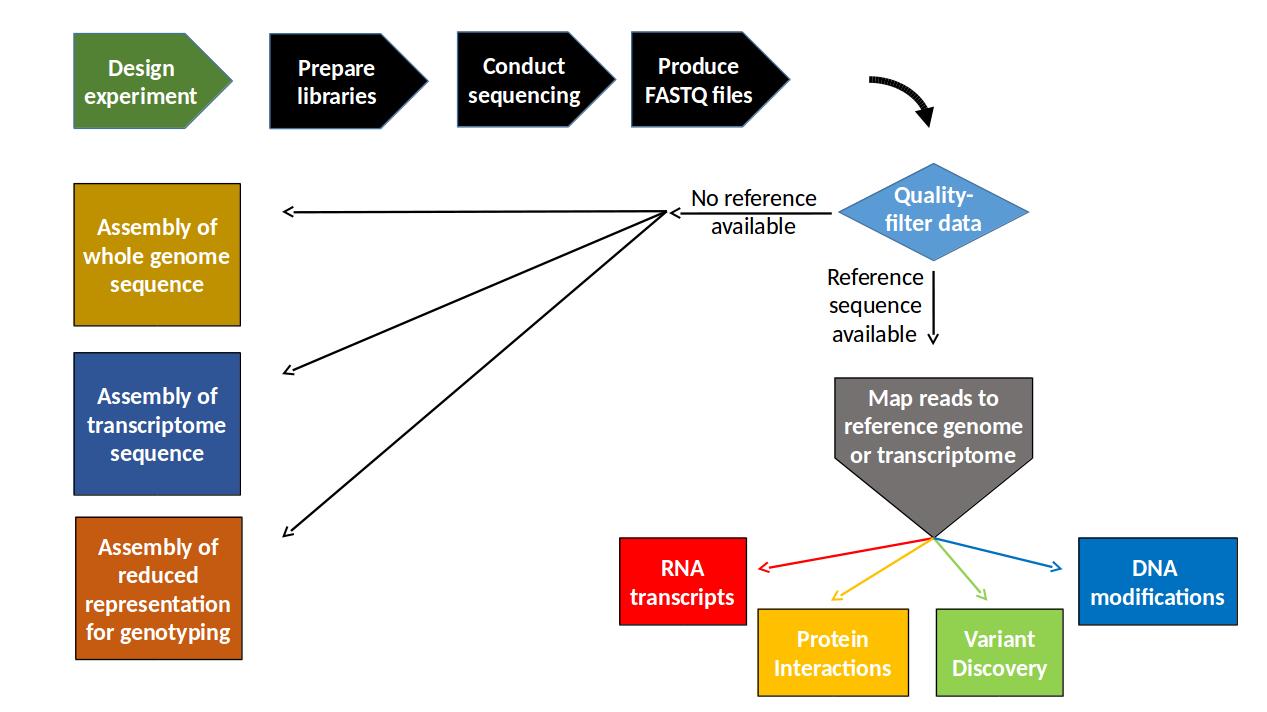
A flow-chart overview of DNA sequencing experiments¶
Last modified 3 January 2022. Edits by Ross Whetten, Will Kohlway, & Maria Adonay.